Almost no-one coming to computational neuroscience has a degree in computational neuroscience: most have degrees in biology, physics, mathematics or AI. Therefore, I often receive the question what good introductory material is. Here, I compiled a list of some introductory material I like. And don’t get scared, here’s a nice ’ten rules’ paper on how to structure your project, or any multi-disciplinary project. And here are some answers to FAQs about computational neuroscience by Kanaka Rajan.
NB There are several programs you can follow if you want to go into (computational) neuroscience. This is a list of all the Neuro-based MSc programs in Nijmegen. From September 2026, a new BSc program in neuroscience will start!
Videos
Introductory
General introduction into computational neuroscience by the Bernstein Centers in Germany:
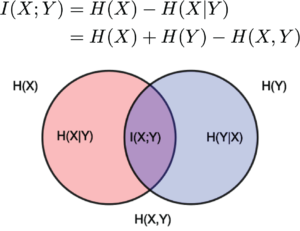
Explanation of Aeon Magazine about the importance of the definition of information by Claude Shannon:
Academic

For some general inspiration, I really like the ‘Growing up in Science‘ series!
Spike coding networks
Here are some videos explaining the efficient spike coding networks we use as a model, by Christian Machens and Sander Keemink:
-
- https://www.youtube.com/watch?v=-1dpfd1mS_E — Christian Machens in a tutorial on how the networks work
- https://www.youtube.com/watch?v=s6xxDf3i3Xo — Christian Machens on more recent developments
- https://www.youtube.com/watch?v=eIUb3dgOHUQ — Sander Keemink on energy efficiency
- https://www.youtube.com/watch?v=Z0V2Xxjpwjg — Michele Nardin on doing general computations with these networks
Literature
Lists made by others
- Dan Goodman makes a nice list of freely available resources. He made a
- The Open Source Brain project also keeps track of the recent and available books
- This GitHub list contains books, data, courses and much more!
- This Comp Neuro website keeps track of labs and resources
My personal preferences
- Scholarpedia: peer reviewed short explanations about a plethora of topics, often by the original authors themselves!
- Dayan, P., & Abbott, L. F. (2001). Theoretical Neuroscience: Computational and Mathematical Modeling of Neural Systems. MIT Press.
- Gerstner, W., Kistler, W. M., Naud, R., & Paninski, L. (2014). Neuronal Dynamics (freely available online)
- Izhikevich, E. M. (2007). Dynamical Systems in Neuroscience: The Geometry of Excitability and Bursting. MIT Press. (online)
- Rieke, F., Warland, D., de Ruyter van Steveninck, R. R., & Bialek, W. (1997). Spikes: Exploring the neural code. In T. J. Sejnowski & T. A. Poggio (Eds.), Computational neuroscience. MIT Press.
-
Miller, P. (2018). An introductory course in computational neuroscience. The MIT Press.
- Lindsay, G. (2021). MODELS OF THE MIND: How physics, engineering and mathematics have shaped our understanding of the brain. BLOOMSBURY SIGMA.
- Robert Rosenbaum recently made his book ‘Modeling Neural Circuits Made Simple’ including all the code available online, a great introduction to computational neuroscience!
Online tutorials
Linear Algebra and more essential mathematics
In many Dutch universities, BSc curricula in Biology do not include linear algebra (i.e. doing calculations with vectors/matrices) or Fourier analysis (i.e. looking what frequency oscillations are in a signal). Here are some good introductory courses
- Mathematical Tools for Neuroscience by Ella Batty (includes exercises in Python)
- Mathematical Tools for Neural and Cognitive Science by Mike Landy and Eero Simoncelli (includes exercises in Matlab)
- Convolutions can be confusing when you start using them (or even when you have used them for years). This blog is great for getting an intuition about them!
- I have heard good stories about the 3Blue1Brown channel on youtube explaining some general maths topics
- Similarly, this Youtube Playlist explains essential calculus ‘like something you yourself could have discovered’
- The Khan Academy is a good starting point for almost any mathematical topic
- Explained Visually has a few good explanation for the more visually oriented among us
- Online material about mathematical modelling in Biology from the course ‘Modeling Life’ by Alan Garfunkel
Modelling in Neuroscience
- Brian is a great resource for starting making models, and it also has some tutorials available. Great to learn both brain simulations and programming in Python!
- The Brain Dynamics Toolbox is a great tool for dynamical systems
- For an introduction in programming, there are many resources, Datacamp is a good one.
- Kaggle is another good resource.
- This Coursera course by Adrienne Fairhall and Rajesh Rao is a great introduction to computational neuroscience!
- The first few chapters of the book ‘The programmers brain’ by Felienne Hermans on how to learn how to program you can read for free!
- The Neuromatch Academy has put all of its materials online
- Dan Goodman gave this Cosyne tutorial and put his material online: Spiking Neural Network Models in Neuroscience
- Jack Terwilliger writes about basic concepts in theoretical neuroscience in his blog.
Machine learning
- A great ‘normal language’ explanation: vas3k
- A great starting kit in Python: scikit-learn
- A great set of video lectures introducting machine learning
- And a follow-up: video lectures on neural data science
- The Neuromatch Academy has put all of its materials online
Best Practices for writing code and organising data
Some random tips from myself.
- Separate the simulation (i.e. the code that generates data) and the analysis. So basically, make a folder with code that does the simulations, a folder with results from these simulations (i.e. ‘raw data’) and a separate folder with analysis code and results
- Store your simulations settings, your parameters, with your data, so that you can always see what parameters were used for a simulation
- Generate data-filenames automatically, in a logical way (i.e. the type of network, the data, maybe a parameter value)
- In your code: choose local names for variables and parameters (i.e. not ‘x’and ‘y’ but ‘membrane_potential’ and ‘input_current’)
- Comment! Explain what you do and why! And make README files on how to use your code. Start a function with a summary, and what goes in and what goes out.
- Use functions and/or classes/objects to organize your code, no long scripts please! Rule of thumb: more than 20 or so lines of code means a function.
- Make a single ‘settings’ structure or library with your chosen variables and parameters, that you both pass to your simulation function and store with your raw data (see 2): this way you can easily add new parameters if you have to, without having to rewrite your functions
- Use version control! Otherwise, you will one day ‘break’ your code and not be able to put it back in working order.
- Do NOT use global variables, ever!
- Do NOT comment out parts of your code if you want to change a setting.
- Do NOT copy-paste pieces of code: if you want to do that you have to create a function
- Put ‘sanity checks’ in your code, and create logical error messages.
- Use the debugging tools that your system provides. For Matlab: try…catch, keyboard, for Python ipdb tools.
- For Python: use Anaconda
- A tip for supervisors: let a new student start with existing code and let him/her update the code with comments. This way, the student will learn what makes a code easy or hard to understand, and the code itself gets improved!
- Another tip for supervisors: (peer) code review sessions (see also some best practices here, here)
Tools and references
This is not an exhaustive list, others have thought more deeply about this:
- Riquelme, J. L., & Gjorgjieva, J. (2021). Towards readable code in neuroscience. Nature Reviews Neuroscience. https://doi.org/10.1038/s41583-021-00450-y
- Carey, M. A., & Papin, J. A. (2018). Ten simple rules for biologists learning to program. PLOS Computational Biology, 14(1), e1005871. https://doi.org/10.1371/journal.pcbi.1005871
-
Vable, A. M., Diehl, S. F., & Glymour, M. M. (2021). Code Review as a Simple Trick to Enhance Reproducibility, Accelerate Learning, and Improve the Quality of Your Team’s Research. American Journal of Epidemiology. https://doi.org/10.1093/aje/kwab092
And some helpful tools and courses:
- Software Carpentry
- Data Carpentry
- NeuralEnsemble (including the ‘Sumatra’ tool for organising simulations)
- Another list of best practices by OCNS Software Special Interest Group
Offline
Summer Schools
As a starting PhD student or a post-doc coming from a different field, a summer school is a great way to learn the most recent advances of the field, and get to know people. I can recommend the following ones:
- OIST Computational Neuroscience Course, Okinawa, Japan
- MBL Woods Hole Methods in Computational Neuroscience
- Cajal Training in Computational Neuroscience (different locations in Europe)
- Mathematical Methods in Computational Neuroscience, Norway
- Simons Computational Neuroscience Imbizo, South Africa
- The online Neuromatch Academy is a great alternative in lockdown times
- The online SNUFA conference focuses on machine learning in spiking networks
Courses in Dutch / in the Netherlands
Physical courses in the Netherlands: Marieke van Vugt Mario Negrello, Jorge Mejias and me try to keep track of all computational neuroscience university courses in the Netherlands. Please take a look here!
For some introductory slides in Dutch, here are some lectures slides about modelling of neurons and networks (1, 2, 3) an introduction to decoding, some introductory slides about (un)supervised learning (1, 2) and reinforcement learning (1 ,2).